Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Ishida, Hisashi; Go, Nobuhiro
no journal, ,
no abstracts in English
Kono, Hidetoshi; Kumar, S.*; Ahmad, S.*; Ara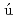 zo-Bravo, M. J.*; Fujii, Satoshi*; Go, Nobuhiro; Sarai, Akinori*
zo-Bravo, M. J.*; Fujii, Satoshi*; Go, Nobuhiro; Sarai, Akinori*
no journal, ,
no abstracts in English
Kim, T. P. O.; Yura, Kei; Go, Nobuhiro
no journal, ,
Protein-RNA interactions play essential roles in a number of regulatory mechanisms of gene expression such as RNA splicing, transport and translation. Computational analyses of protein-DNA interface have been carried out a lot, while those of protein-RNA interface are rare. As the number of available protein-RNA complexes have increased, it is now possible to examine protein-RNA interactions based on 3D structures. We carried out computational analyses of protein-RNA complexes retrieved from PDB. The interface residue propensity was used as a parameter to predict protein-RNA interface. The accuracy of the prediction method was tested by jackknife method, which attained 60% on average. The prediction method was applied to RNA-binding proteins with known 3D structures (mRNA export factors TAP and Mex67).
Matsumoto, Atsushi
no journal, ,
We take advantage of a new approach to study the low frequency normal modes of DNA fragments of a few hundred base pairs in terms of the constituent base-pair steps. The advantage of this approach is that calculations for relatively long DNA fragments are very fast and yet the sequence-dependent local structure and flexibility of the polymer is considered. Initial application of the approach to linear DNA revealed the relationship between global deformations, such as bending and twisting, and sequential variations of the base-pair step parameters. Subsequent extension of the calculations to circular DNA showed remarkable agreement with the theoretically predicted dynamical properties of a circular DNA modeled as an ideal, inextensible elastic rod and revealed new insights into the global configurational rearrangement of the circle to the figure-8 form. We now study more realistically modeled circular DNA and examine the effects of local conformational properties, such as the intrinsic bending, twisting, and flexibility of base-pair steps, on global structure and dynamics. We focus on the intrinsic flexibility of the p53 response element DNA, which has been previously studied in ring closure experiments. We will discuss the origin of this flexibility in terms of local conformational properties.
Nakagawa, Hiroshi
no journal, ,
Incoherent neutron scattering is one of the effective methods to investigate picosecond and nanosecond time scale dynamics of protein. In order to understand the dynamical properties of protein, the effects of hydration and temperature on the dynamics of Staphylococcal nuclease were examined by incoherent inelastic neutron scattering. At the cryogenic temperature the protein boson peak was observed in the low frequency region, and the hydration shifts the peak frequency to higher frequency. This indicates that the vibrational low frequency mode is closely coupled with the solvent. The anomalous decrease in the Debye-Waller factor at higher temperature corresponds to an increase of the mean-square displacement, which is called dynamical transition. This is accompanied by the appearance of a quasielastic scattering. The quasielastic scattering comes from anharmonic motions such as relaxation and/or stochastic motions. Hydration dependent and independent dynamical transition were observed. Analysis of quasielastic scattering could elucidate the dynamical features of the transition, which may be essential for protein function.
Nakagawa, Hiroshi; Jochi, Yasumasa*; Kataoka, Mikio*; Go, Nobuhiro
no journal, ,
no abstracts in English
Hikuchi, Mariko; Ishida, Hisashi; Kitao, Akio*; Yamagata, Yuriko*; Go, Nobuhiro
no journal, ,
no abstracts in English
Yura, Kei
no journal, ,
no abstracts in English
Fujiwara, Satoru; Matsumoto, Fumiko*; Takahashi, Nobuaki
no journal, ,
no abstracts in English